Plugin Marketplace — Design Document
Overview
The Plugin Marketplace provides an extensible architecture for integrating third-party computational tools into the ChemLib Platform. Plugins can be used as filter nodes in screening pipelines, as standalone tools, or as alternative implementations for existing platform features (e.g., alternative docking engines).
The system is designed in three phases:
1. Phase 1 (MVP): Built-in plugins only, shipped with ChemLib
2. Phase 2: pip-installable plugin packages discovered via Python entry points
3. Phase 3: Web-based marketplace with community contributions
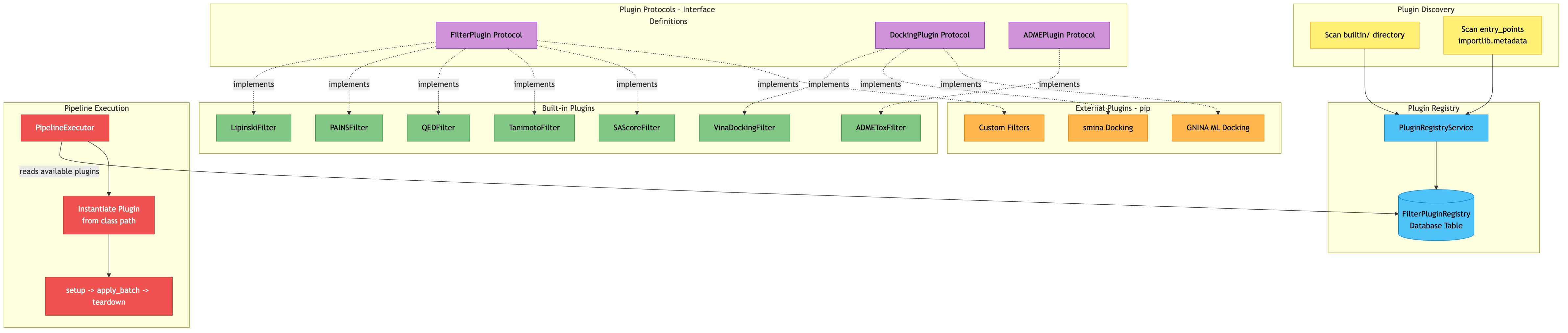
Architecture
Click diagram to zoom and pan:
┌─────────────────────────────────────────────────────────────────────────┐
│ PLUGIN ARCHITECTURE │
│ │
│ ┌──────────────────┐ ┌──────────────────┐ ┌────────────────────┐│
│ │ Built-in Plugins │ │ Entry Point │ │ Future: ││
│ │ (chemlib.plugins │ │ Plugins │ │ Web Marketplace ││
│ │ .builtin.*) │ │ (pip packages) │ │ (download + reg) ││
│ └────────┬─────────┘ └────────┬─────────┘ └────────┬───────────┘│
│ │ │ │ │
│ ▼ ▼ ▼ │
│ ┌─────────────────────────────────────────────────────────────────┐ │
│ │ Plugin Registry │ │
│ │ chemlib/plugins/registry.py │ │
│ │ │ │
│ │ - Discovers plugins (built-in + entry points) │ │
│ │ - Validates Protocol compliance │ │
│ │ - Registers in FilterPluginRegistry table │ │
│ │ - Instantiates plugin objects on demand │ │
│ └───────────────────────────┬─────────────────────────────────────┘ │
│ │ │
│ ▼ │
│ ┌─────────────────────────────────────────────────────────────────┐ │
│ │ FilterPluginRegistry (Database) │ │
│ │ Stores: name, category, description, config_schema, class path │ │
│ └───────────────────────────┬─────────────────────────────────────┘ │
│ │ │
│ ┌───────────────────┼───────────────────┐ │
│ ▼ ▼ ▼ │
│ ┌────────────────┐ ┌────────────────┐ ┌────────────────────┐ │
│ │ Pipeline │ │ Standalone │ │ Marketplace UI │ │
│ │ Builder │ │ Tool Usage │ │ (browse, config) │ │
│ │ (DAG nodes) │ │ (API calls) │ │ │ │
│ └────────────────┘ └────────────────┘ └────────────────────┘ │
└─────────────────────────────────────────────────────────────────────────┘
Plugin Lifecycle
1. DISCOVERY
Plugin exists as:
- Python class in chemlib/plugins/builtin/
- OR: installed pip package with entry point "chemlib.plugins"
2. REGISTRATION
PluginRegistryService.register_all() on app startup:
- Scan built-in plugins (import all classes)
- Scan entry points (pkg_resources / importlib.metadata)
- For each: validate Protocol compliance, extract metadata
- Upsert into FilterPluginRegistry table
3. CONFIGURATION
User selects a plugin in the Pipeline Builder or Marketplace UI:
- config_schema (JSON Schema) is fetched from DB
- UI renders a form from the schema
- User fills in parameters (thresholds, references, etc.)
4. EXECUTION
Plugin is used in a pipeline run:
- PipelineExecutor instantiates the plugin class
- Calls setup() with config and context
- Calls apply() or apply_batch() for each compound
- Calls teardown() after all compounds processed
5. DEACTIVATION (optional)
Admin can set is_active=False in the registry to disable a plugin.
Plugin Protocol Definitions
All plugin types are defined as Python Protocol classes in chemlib/plugins/protocols.py. This allows structural subtyping — any class that implements the required methods and properties is valid, without needing to inherit from a base class.
FilterPlugin (Primary — used in screening pipelines)
# Defined in SCREENING_PIPELINE.md — reproduced here for reference
@runtime_checkable
class FilterPlugin(Protocol):
@property
def name(self) -> str: ...
@property
def display_name(self) -> str: ...
@property
def category(self) -> str: ...
@property
def description(self) -> str: ...
@property
def config_schema(self) -> dict: ...
@property
def estimated_time_per_compound(self) -> str: ...
async def apply(self, mol: Chem.Mol, config: dict, context: PipelineContext) -> FilterResult: ...
async def apply_batch(self, mols: list[Chem.Mol], config: dict, context: PipelineContext) -> list[FilterResult]: ...
async def setup(self, config: dict, context: PipelineContext) -> None: ...
async def teardown(self) -> None: ...
DockingPlugin (Alternative docking engines)
@runtime_checkable
class DockingPlugin(Protocol):
"""Protocol for alternative docking engines (smina, GNINA, etc.)."""
@property
def name(self) -> str:
"""Unique plugin name (e.g., 'smina_docking')."""
...
@property
def display_name(self) -> str:
"""Human-readable name (e.g., 'smina Docking Engine')."""
...
@property
def description(self) -> str: ...
@property
def config_schema(self) -> dict:
"""
JSON Schema for docking parameters.
Example: exhaustiveness, scoring function, flexible residues, etc.
"""
...
@property
def supports_flexible_residues(self) -> bool:
"""Whether this engine supports flexible receptor residues."""
...
async def prepare_receptor(self, pdb_data: str, config: dict) -> str:
"""
Prepare receptor for this docking engine.
Returns: engine-specific receptor format (e.g., PDBQT, MOL2).
"""
...
async def prepare_ligand(self, smiles: str, config: dict) -> str:
"""
Prepare ligand for this docking engine.
Returns: engine-specific ligand format.
"""
...
async def dock(
self,
receptor_data: str,
ligand_data: str,
center: tuple[float, float, float],
box_size: tuple[float, float, float],
config: dict
) -> DockingPluginResult:
"""
Run docking.
Returns: DockingPluginResult with scores and poses.
"""
...
@dataclass
class DockingPluginResult:
scores: list[float] # kcal/mol per pose, sorted best to worst
poses_data: str # All poses in engine-native format
poses_pdb: list[str] # Poses converted to PDB format for visualization
best_score: float
metadata: dict = field(default_factory=dict)
@runtime_checkable
class ADMEPlugin(Protocol):
"""Protocol for ADME/toxicity prediction tools."""
@property
def name(self) -> str: ...
@property
def display_name(self) -> str: ...
@property
def description(self) -> str: ...
@property
def config_schema(self) -> dict: ...
@property
def predicted_properties(self) -> list[str]:
"""
List of ADME/Tox properties this plugin predicts.
Example: ["solubility", "bbb_penetration", "cyp_inhibition", "herg_liability"]
"""
...
async def predict(self, mol: Chem.Mol, config: dict) -> ADMEPrediction:
"""
Predict ADME/Tox properties for a single molecule.
"""
...
async def predict_batch(self, mols: list[Chem.Mol], config: dict) -> list[ADMEPrediction]:
"""Batch prediction."""
...
@dataclass
class ADMEPrediction:
properties: dict[str, Any] # {"solubility": -3.2, "bbb": True, "herg": "low_risk"}
confidence: dict[str, float] | None = None # {"solubility": 0.85, "bbb": 0.72}
metadata: dict = field(default_factory=dict)
VisualizationPlugin (Custom visualization renderers)
@runtime_checkable
class VisualizationPlugin(Protocol):
"""Protocol for custom visualization components."""
@property
def name(self) -> str: ...
@property
def display_name(self) -> str: ...
@property
def description(self) -> str: ...
@property
def output_format(self) -> str:
"""Output format: 'html', 'svg', 'png', 'json'."""
...
async def render(self, data: dict, config: dict) -> str:
"""
Render visualization from input data.
Returns: HTML string, SVG string, base64-encoded PNG, or JSON.
"""
...
ExternalServicePlugin (REST API wrappers)
@runtime_checkable
class ExternalServicePlugin(Protocol):
"""Protocol for external REST API integrations."""
@property
def name(self) -> str: ...
@property
def display_name(self) -> str: ...
@property
def description(self) -> str: ...
@property
def config_schema(self) -> dict: ...
@property
def base_url(self) -> str: ...
@property
def rate_limit(self) -> int:
"""Maximum requests per second to this service."""
...
async def query(self, mol: Chem.Mol, config: dict) -> dict:
"""
Query external API for a single molecule.
Returns: API response parsed as dict.
"""
...
async def health_check(self) -> bool:
"""Check if the external service is reachable."""
...
Plugin Registry
chemlib/plugins/registry.py
"""Plugin discovery, validation, and registration."""
from __future__ import annotations
import importlib
import importlib.metadata
import logging
from typing import Type, Any
from sqlalchemy import select
from sqlalchemy.ext.asyncio import AsyncSession
from chemlib.models.plugin import FilterPluginRegistry
from chemlib.plugins.protocols import FilterPlugin, DockingPlugin, ADMEPlugin
logger = logging.getLogger(__name__)
# Entry point group names
FILTER_PLUGIN_GROUP = "chemlib.plugins.filter"
DOCKING_PLUGIN_GROUP = "chemlib.plugins.docking"
ADME_PLUGIN_GROUP = "chemlib.plugins.adme"
class PluginRegistryService:
"""Manages plugin discovery, registration, and instantiation."""
# In-memory cache of instantiated plugins
_plugin_cache: dict[str, Any] = {}
async def discover_and_register_all(self, db: AsyncSession) -> int:
"""
Discover all plugins (built-in + entry points) and register them.
Called once at app startup.
Returns: number of plugins registered.
"""
count = 0
count += await self._register_builtin_plugins(db)
count += await self._register_entrypoint_plugins(db)
await db.commit()
logger.info(f"Registered {count} plugins total")
return count
async def _register_builtin_plugins(self, db: AsyncSession) -> int:
"""
Import and register all built-in plugin classes from
chemlib.plugins.builtin.* modules.
"""
builtin_modules = [
"chemlib.plugins.builtin.property_filters",
"chemlib.plugins.builtin.similarity_filters",
"chemlib.plugins.builtin.adme_filters",
"chemlib.plugins.builtin.docking_filter",
"chemlib.plugins.builtin.external_filters",
]
count = 0
for module_path in builtin_modules:
try:
module = importlib.import_module(module_path)
# Find all classes in the module that implement FilterPlugin
for attr_name in dir(module):
attr = getattr(module, attr_name)
if (
isinstance(attr, type)
and attr_name != "FilterPlugin"
and hasattr(attr, 'name')
and hasattr(attr, 'apply')
):
plugin = attr()
if isinstance(plugin, FilterPlugin):
await self._register_plugin(db, plugin, module_path, is_builtin=True)
count += 1
except ImportError as e:
logger.warning(f"Could not import built-in module {module_path}: {e}")
return count
async def _register_entrypoint_plugins(self, db: AsyncSession) -> int:
"""
Discover plugins via Python entry points.
Third-party packages register their plugins in pyproject.toml:
[project.entry-points."chemlib.plugins.filter"]
my_filter = "my_package.filters:MyCustomFilter"
"""
count = 0
for group in [FILTER_PLUGIN_GROUP, DOCKING_PLUGIN_GROUP, ADME_PLUGIN_GROUP]:
try:
entry_points = importlib.metadata.entry_points(group=group)
for ep in entry_points:
try:
plugin_class = ep.load()
plugin = plugin_class()
module_path = f"{ep.value}"
await self._register_plugin(db, plugin, module_path, is_builtin=False)
count += 1
logger.info(f"Registered entry point plugin: {ep.name} from {ep.value}")
except Exception as e:
logger.error(f"Failed to load entry point {ep.name}: {e}")
except Exception as e:
logger.warning(f"Could not read entry points for {group}: {e}")
return count
async def _register_plugin(
self, db: AsyncSession,
plugin: FilterPlugin,
module_path: str,
is_builtin: bool
) -> None:
"""
Upsert a plugin into the FilterPluginRegistry table.
If the plugin already exists (by name), update its metadata.
Otherwise, insert a new record.
"""
result = await db.execute(
select(FilterPluginRegistry).where(
FilterPluginRegistry.name == plugin.name
)
)
existing = result.scalar_one_or_none()
class_path = f"{module_path}.{plugin.__class__.__name__}"
if existing:
existing.display_name = plugin.display_name
existing.category = plugin.category
existing.description = plugin.description
existing.plugin_class = class_path
existing.config_schema = plugin.config_schema
existing.estimated_time = plugin.estimated_time_per_compound
existing.is_builtin = is_builtin
else:
db.add(FilterPluginRegistry(
name=plugin.name,
display_name=plugin.display_name,
category=plugin.category,
description=plugin.description,
plugin_class=class_path,
config_schema=plugin.config_schema,
estimated_time=plugin.estimated_time_per_compound,
is_builtin=is_builtin,
is_active=True,
version="1.0.0",
))
async def get_plugin_instance(self, plugin_name: str) -> FilterPlugin:
"""
Get an instantiated plugin by name.
Uses in-memory cache to avoid re-importing.
Raises: PluginNotFoundError if not registered.
"""
if plugin_name in self._plugin_cache:
return self._plugin_cache[plugin_name]
# Look up in DB
# Import the class, instantiate, cache, return
...
async def list_plugins(
self, db: AsyncSession,
category: str | None = None,
active_only: bool = True
) -> list[FilterPluginRegistry]:
"""List registered plugins with optional category filter."""
query = select(FilterPluginRegistry)
if category:
query = query.where(FilterPluginRegistry.category == category)
if active_only:
query = query.where(FilterPluginRegistry.is_active == True)
result = await db.execute(query.order_by(FilterPluginRegistry.category, FilterPluginRegistry.name))
return result.scalars().all()
async def get_plugin_info(
self, db: AsyncSession, plugin_name: str
) -> FilterPluginRegistry | None:
"""Get plugin metadata from the registry."""
result = await db.execute(
select(FilterPluginRegistry).where(FilterPluginRegistry.name == plugin_name)
)
return result.scalar_one_or_none()
async def set_active(
self, db: AsyncSession, plugin_name: str, active: bool
) -> bool:
"""Enable or disable a plugin."""
result = await db.execute(
select(FilterPluginRegistry).where(FilterPluginRegistry.name == plugin_name)
)
plugin = result.scalar_one_or_none()
if plugin:
plugin.is_active = active
await db.commit()
return True
return False
Built-in Plugins (Phase 1 — Seed the Marketplace)
Complete List by Category
Docking
| Plugin |
Class |
Description |
External Dep |
| AutoDock Vina |
VinaDockingFilter |
Dock compound against protein binding site |
vina, meeko |
| smina (optional) |
SminaDockingFilter |
Vina-based engine with custom scoring functions |
smina binary |
| GNINA (optional) |
GninaDockingFilter |
CNN-scored molecular docking |
gnina binary |
Pocket Detection
| Plugin |
Class |
Description |
External Dep |
| Fpocket |
FpocketDetector |
Detect druggable binding pockets |
fpocket binary |
Protein Preparation
| Plugin |
Class |
Description |
External Dep |
| PDBFixer |
PDBFixerPrep |
Fix missing atoms/residues, add H |
pdbfixer |
| Plugin |
Class |
Description |
External Dep |
| Open Babel |
OpenBabelConverter |
Convert between PDB, PDBQT, MOL2, SDF |
openbabel-wheel |
Interaction Analysis
| Plugin |
Class |
Description |
External Dep |
| PLIP |
PLIPAnalyzer |
Protein-ligand interaction profiler |
plip |
Property Filters
| Plugin |
Class |
Description |
External Dep |
| Lipinski RO5 |
LipinskiFilter |
MW, LogP, HBD, HBA thresholds |
RDKit |
| Veber Rules |
VeberFilter |
TPSA, RotBonds |
RDKit |
| Ghose Filter |
GhoseFilter |
MW, LogP, atoms, MR ranges |
RDKit |
| Egan Filter |
EganFilter |
TPSA, LogP |
RDKit |
| Muegge Filter |
MueggeFilter |
Multi-parameter pharmaceutical filter |
RDKit |
| PAINS |
PAINSFilter |
Pan-assay interference substructures |
RDKit |
| Brenk |
BrenkFilter |
Structural alerts |
RDKit |
| QED Threshold |
QEDThresholdFilter |
Drug-likeness score threshold |
RDKit |
| SA Score |
SAScoreFilter |
Synthetic accessibility threshold |
RDKit |
| MW Range |
MWRangeFilter |
Molecular weight range |
RDKit |
| LogP Range |
LogPRangeFilter |
Lipophilicity range |
RDKit |
| TPSA Range |
TPSARangeFilter |
Polar surface area range |
RDKit |
| HBD Max |
HBDMaxFilter |
Max hydrogen bond donors |
RDKit |
| HBA Max |
HBAMaxFilter |
Max hydrogen bond acceptors |
RDKit |
| RotBonds Max |
RotBondsMaxFilter |
Max rotatable bonds |
RDKit |
Similarity Filters
| Plugin |
Class |
Description |
External Dep |
| Morgan FP Tanimoto |
TanimotoSimilarityFilter |
Similarity to reference compound |
RDKit |
| Substructure Match |
SubstructureMatchFilter |
Contains SMARTS substructure |
RDKit |
| MACCS Similarity |
MACCSSimilarityFilter |
MACCS keys similarity |
RDKit |
ADME Prediction
| Plugin |
Class |
Description |
External Dep |
| ESOL Solubility |
ESOLSolubilityFilter |
Delaney aqueous solubility prediction |
RDKit |
| Simple BBB |
BBBRuleFilter |
Blood-brain barrier rule |
RDKit |
| Simple hERG |
hERGRuleFilter |
hERG cardiac liability rule |
RDKit |
| Rule of Three |
RuleOfThreeFilter |
Fragment-likeness filter |
RDKit |
| DeepChem ADME (optional) |
DeepChemADMEFilter |
ML-based ADME predictions |
deepchem |
External APIs
| Plugin |
Class |
Description |
External Dep |
| ADMETlab 2.0 |
ADMETlabFilter |
Online ADME/Tox prediction |
HTTP client |
| ChEMBL Lookup |
ChEMBLLookupPlugin |
Check if compound exists in ChEMBL |
HTTP client |
Visualization
| Plugin |
Class |
Description |
External Dep |
| 3Dmol.js Viewer |
ThreeDmolViewer |
Interactive 3D molecular viewer |
3Dmol.js (JS) |
| pyMSAviz |
PyMSAvizRenderer |
Static alignment image generator |
pymsaviz |
Plugin Configuration
Each plugin exposes a config_schema property that returns a JSON Schema object. The UI renders a configuration form automatically from this schema.
| JSON Schema Type |
UI Element |
"type": "string" |
Text input |
"type": "integer" |
Number input (step=1) |
"type": "number" |
Number input (step=0.1) |
"type": "boolean" |
Checkbox |
"enum": [...] |
Dropdown select |
"type": "string" with "format": "smiles" |
SMILES input with structure preview |
"minimum", "maximum" |
Range slider or bounded input |
Example Config Schemas
Lipinski Filter
{
"type": "object",
"properties": {
"allow_violations": {
"type": "integer",
"minimum": 0,
"maximum": 2,
"default": 0,
"title": "Allowed Violations",
"description": "Number of Lipinski rule violations to allow (0 = strict, 2 = lenient)"
}
}
}
Tanimoto Similarity Filter
{
"type": "object",
"properties": {
"reference_smiles": {
"type": "string",
"format": "smiles",
"title": "Reference Compound SMILES",
"description": "SMILES of the compound to compare against"
},
"threshold": {
"type": "number",
"minimum": 0.0,
"maximum": 1.0,
"default": 0.5,
"title": "Minimum Tanimoto Similarity",
"description": "Compounds with Tanimoto >= threshold pass the filter"
},
"radius": {
"type": "integer",
"minimum": 1,
"maximum": 4,
"default": 2,
"title": "Morgan FP Radius",
"description": "Radius for Morgan/ECFP fingerprint generation"
},
"nbits": {
"type": "integer",
"enum": [1024, 2048, 4096],
"default": 2048,
"title": "Fingerprint Bits",
"description": "Size of the bit vector for fingerprint"
}
},
"required": ["reference_smiles"]
}
AutoDock Vina Docking
{
"type": "object",
"properties": {
"binding_site_id": {
"type": "integer",
"title": "Binding Site",
"description": "Select the binding site to dock against"
},
"exhaustiveness": {
"type": "integer",
"minimum": 8,
"maximum": 128,
"default": 32,
"title": "Exhaustiveness",
"description": "Search thoroughness. Higher values are slower but more accurate."
},
"num_poses": {
"type": "integer",
"minimum": 1,
"maximum": 50,
"default": 10,
"title": "Number of Poses",
"description": "Maximum number of binding poses to generate"
},
"score_threshold": {
"type": "number",
"maximum": 0,
"default": -6.0,
"title": "Score Threshold (kcal/mol)",
"description": "Compounds scoring better (more negative) than this pass"
},
"energy_range": {
"type": "number",
"minimum": 1.0,
"maximum": 10.0,
"default": 3.0,
"title": "Energy Range (kcal/mol)",
"description": "Maximum energy difference between best and worst pose"
}
},
"required": ["binding_site_id"]
}
// chemlib/static/js/plugin_config_form.js
/**
* Render a JSON Schema as an HTML form.
* Used in the Pipeline Builder (node config panel) and Marketplace (plugin config).
*/
function renderConfigForm(schema, containerId, currentValues = {}) {
const container = document.getElementById(containerId);
container.innerHTML = '';
if (!schema || !schema.properties) return;
for (const [key, prop] of Object.entries(schema.properties)) {
const div = document.createElement('div');
div.className = 'config-field';
const label = document.createElement('label');
label.textContent = prop.title || key;
label.setAttribute('for', `config_${key}`);
div.appendChild(label);
if (prop.description) {
const hint = document.createElement('small');
hint.textContent = prop.description;
div.appendChild(hint);
}
let input;
if (prop.enum) {
// Dropdown select
input = document.createElement('select');
for (const val of prop.enum) {
const option = document.createElement('option');
option.value = val;
option.textContent = val;
if (val === (currentValues[key] ?? prop.default)) option.selected = true;
input.appendChild(option);
}
} else if (prop.type === 'boolean') {
// Checkbox
input = document.createElement('input');
input.type = 'checkbox';
input.checked = currentValues[key] ?? prop.default ?? false;
} else if (prop.type === 'integer' || prop.type === 'number') {
// Number input
input = document.createElement('input');
input.type = 'number';
input.step = prop.type === 'integer' ? '1' : '0.1';
if (prop.minimum !== undefined) input.min = prop.minimum;
if (prop.maximum !== undefined) input.max = prop.maximum;
input.value = currentValues[key] ?? prop.default ?? '';
} else {
// Text input
input = document.createElement('input');
input.type = 'text';
input.value = currentValues[key] ?? prop.default ?? '';
if (prop.format === 'smiles') {
input.placeholder = 'Enter SMILES...';
input.addEventListener('change', () => previewSmiles(input.value, div));
}
}
input.id = `config_${key}`;
input.name = key;
div.appendChild(input);
// Required indicator
if (schema.required && schema.required.includes(key)) {
const req = document.createElement('span');
req.className = 'required';
req.textContent = '*';
label.appendChild(req);
}
container.appendChild(div);
}
}
function getConfigValues(containerId) {
const container = document.getElementById(containerId);
const values = {};
for (const input of container.querySelectorAll('input, select')) {
if (input.type === 'checkbox') {
values[input.name] = input.checked;
} else if (input.type === 'number') {
values[input.name] = parseFloat(input.value);
} else {
values[input.name] = input.value;
}
}
return values;
}
Marketplace UI
Plugin Browser Page (plugin_marketplace.html)
┌─────────────────────────────────────────────────────────────────────────┐
│ Plugin Marketplace [Refresh] │
├─────────────────────────────────────────────────────────────────────────┤
│ │
│ Categories: [All ▾] Search: [________________] [Built-in only: ☑] │
│ │
│ ┌───────────────────────────────────────────────────────────────────┐ │
│ │ PROPERTY FILTERS (15) │ │
│ ├───────────────────────────────────────────────────────────────────┤ │
│ │ │ │
│ │ ┌─────────────────────┐ ┌─────────────────────┐ │ │
│ │ │ Lipinski RO5 │ │ PAINS Filter │ │ │
│ │ │ ─────────────────── │ │ ─────────────────── │ │ │
│ │ │ Filters by Lipinski │ │ Pan-Assay │ │ │
│ │ │ Rule of Five: │ │ Interference │ │ │
│ │ │ MW, LogP, HBD, HBA │ │ Compounds filter │ │ │
│ │ │ │ │ │ │ │
│ │ │ ■ Built-in ● Active│ │ ■ Built-in ● Active│ │ │
│ │ │ ~ms/compound │ │ ~ms/compound │ │ │
│ │ │ │ │ │ │ │
│ │ │ [Configure] [Use] │ │ [Configure] [Use] │ │ │
│ │ └─────────────────────┘ └─────────────────────┘ │ │
│ │ │ │
│ │ ┌─────────────────────┐ ┌─────────────────────┐ │ │
│ │ │ QED Score │ │ SA Score │ │ │
│ │ │ ─────────────────── │ │ ─────────────────── │ │ │
│ │ │ Drug-likeness score │ │ Synthetic │ │ │
│ │ │ threshold filter │ │ accessibility score │ │ │
│ │ │ ... │ │ ... │ │ │
│ │ └─────────────────────┘ └─────────────────────┘ │ │
│ └───────────────────────────────────────────────────────────────────┘ │
│ │
│ ┌───────────────────────────────────────────────────────────────────┐ │
│ │ DOCKING (1-3) │ │
│ ├───────────────────────────────────────────────────────────────────┤ │
│ │ ┌─────────────────────┐ │ │
│ │ │ AutoDock Vina │ │ │
│ │ │ ─────────────────── │ │ │
│ │ │ Molecular docking │ │ │
│ │ │ against protein │ │ │
│ │ │ binding sites │ │ │
│ │ │ │ │ │
│ │ │ ■ Built-in ● Active│ │ │
│ │ │ ~min/compound │ │ │
│ │ │ │ │ │
│ │ │ [Configure] [Use] │ │ │
│ │ └─────────────────────┘ │ │
│ └───────────────────────────────────────────────────────────────────┘ │
│ │
│ ┌───────────────────────────────────────────────────────────────────┐ │
│ │ SIMILARITY (3) │ │
│ │ ... │ │
│ └───────────────────────────────────────────────────────────────────┘ │
│ │
│ ┌───────────────────────────────────────────────────────────────────┐ │
│ │ ADME (4-5) │ │
│ │ ... │ │
│ └───────────────────────────────────────────────────────────────────┘ │
│ │
│ ┌───────────────────────────────────────────────────────────────────┐ │
│ │ EXTERNAL (2) │ │
│ │ ... │ │
│ └───────────────────────────────────────────────────────────────────┘ │
└─────────────────────────────────────────────────────────────────────────┘
Interactions:
- [Configure]: Opens a modal with the auto-generated config form from config_schema
- [Use in Pipeline]: Opens the Pipeline Builder with this plugin pre-added as a node
- Category filter: Show only plugins from a specific category
- Active toggle: Admin can enable/disable plugins
Plugin Installation (Phased)
Phase 1: Built-in Only
All plugins ship with ChemLib. No installation needed. The seed_plugins.py script registers them in the database on first run. The PluginRegistryService.discover_and_register_all() method is called at app startup.
Phase 2: pip-installable Packages
Third-party developers create Python packages that define plugins and register them via entry points.
Example third-party plugin package (chemlib-gnina-plugin):
# pyproject.toml of the third-party package
[project]
name = "chemlib-gnina-plugin"
version = "1.0.0"
dependencies = ["chemlib-platform"]
[project.entry-points."chemlib.plugins.filter"]
gnina_docking = "chemlib_gnina.filters:GninaDockingFilter"
# chemlib_gnina/filters.py
from chemlib.plugins.protocols import FilterPlugin, FilterResult, PipelineContext
from rdkit import Chem
class GninaDockingFilter:
@property
def name(self) -> str:
return "gnina_docking"
@property
def display_name(self) -> str:
return "GNINA CNN-Scored Docking"
@property
def category(self) -> str:
return "docking"
@property
def description(self) -> str:
return "Molecular docking with GNINA CNN-based scoring function."
@property
def config_schema(self) -> dict:
return {
"type": "object",
"properties": {
"binding_site_id": {"type": "integer", "title": "Binding Site"},
"cnn_scoring": {
"type": "string",
"enum": ["default2018", "dense", "crossdock_default2018"],
"default": "default2018",
"title": "CNN Scoring Function"
},
"exhaustiveness": {
"type": "integer", "minimum": 8, "maximum": 128,
"default": 32, "title": "Exhaustiveness"
}
},
"required": ["binding_site_id"]
}
@property
def estimated_time_per_compound(self) -> str:
return "min"
async def apply(self, mol: Chem.Mol, config: dict, context: PipelineContext) -> FilterResult:
# Run GNINA docking
...
async def apply_batch(self, mols, config, context):
return [await self.apply(m, config, context) for m in mols]
async def setup(self, config, context):
pass
async def teardown(self):
pass
Installation: pip install chemlib-gnina-plugin, then restart ChemLib. The plugin is automatically discovered via entry points and registered.
Phase 3: Web Marketplace (Future)
Potential features:
- Browse community-submitted plugins on a web page
- One-click install (triggers pip install on the server)
- Version management and update notifications
- Plugin ratings and reviews
- Plugin compatibility checking
This phase is out of scope for initial implementation.
API Endpoints
Plugins — chemlib/api/plugins.py
| Method |
Endpoint |
Description |
Response |
GET |
/api/plugins/ |
List all registered plugins |
list[PluginRegistryResponse] |
GET |
/api/plugins/categories |
List available categories |
list[str] |
GET |
/api/plugins/category/{category} |
List plugins by category |
list[PluginRegistryResponse] |
GET |
/api/plugins/{name} |
Get plugin details + config schema |
PluginRegistryResponse |
PUT |
/api/plugins/{name}/active |
Enable/disable a plugin |
{"name": "...", "is_active": true} |
POST |
/api/plugins/refresh |
Re-scan and register plugins |
{"registered": 25} |
# chemlib/api/plugins.py
router = APIRouter(prefix="/api/plugins", tags=["plugins"])
@router.get("/", response_model=list[PluginRegistryResponse])
async def list_plugins(
category: str | None = None,
active_only: bool = True,
db: AsyncSession = Depends(get_db),
service: PluginRegistryService = Depends(),
):
return await service.list_plugins(db, category=category, active_only=active_only)
@router.get("/categories")
async def list_categories(db: AsyncSession = Depends(get_db)):
result = await db.execute(
select(FilterPluginRegistry.category)
.distinct()
.order_by(FilterPluginRegistry.category)
)
return result.scalars().all()
@router.get("/category/{category}", response_model=list[PluginRegistryResponse])
async def list_plugins_by_category(
category: str,
db: AsyncSession = Depends(get_db),
service: PluginRegistryService = Depends(),
):
return await service.list_plugins(db, category=category)
@router.get("/{name}", response_model=PluginRegistryResponse)
async def get_plugin(
name: str,
db: AsyncSession = Depends(get_db),
service: PluginRegistryService = Depends(),
):
plugin = await service.get_plugin_info(db, name)
if not plugin:
raise HTTPException(404, f"Plugin '{name}' not found")
return plugin
@router.put("/{name}/active")
async def set_plugin_active(
name: str,
active: bool,
db: AsyncSession = Depends(get_db),
service: PluginRegistryService = Depends(),
):
success = await service.set_active(db, name, active)
if not success:
raise HTTPException(404, f"Plugin '{name}' not found")
return {"name": name, "is_active": active}
@router.post("/refresh")
async def refresh_plugins(
db: AsyncSession = Depends(get_db),
service: PluginRegistryService = Depends(),
):
count = await service.discover_and_register_all(db)
return {"registered": count}
Testing Strategy
| Component |
Test Type |
Notes |
| Protocol compliance |
Unit |
Verify built-in plugins satisfy Protocol |
| Plugin registry |
Integration |
Register, list, get, activate/deactivate |
| Config schema validation |
Unit |
Validate config against JSON Schema |
| Form rendering |
Frontend |
Manual/Playwright test of auto-generated forms |
| Entry point discovery |
Integration |
Create mock package with entry point |
| Plugin in pipeline |
Integration |
Run pipeline with built-in filter plugin |
Protocol Compliance Test
import pytest
from chemlib.plugins.protocols import FilterPlugin
from chemlib.plugins.builtin.property_filters import LipinskiFilter, PAINSFilter, QEDThresholdFilter
@pytest.mark.parametrize("plugin_class", [
LipinskiFilter,
PAINSFilter,
QEDThresholdFilter,
# ... all built-in plugins
])
def test_filter_plugin_protocol(plugin_class):
"""Verify all built-in plugins satisfy the FilterPlugin protocol."""
plugin = plugin_class()
assert isinstance(plugin, FilterPlugin)
assert isinstance(plugin.name, str) and len(plugin.name) > 0
assert isinstance(plugin.display_name, str)
assert plugin.category in ("property", "similarity", "docking", "adme", "external")
assert isinstance(plugin.description, str)
assert isinstance(plugin.config_schema, dict)
assert plugin.estimated_time_per_compound in ("ms", "s", "min", "hour")
