ChemLib Platform¶
A Computer-Aided Drug Discovery Platform
ChemLib is a comprehensive drug discovery platform built around fragment-based drug design and virtual screening. It provides an integrated environment for managing chemical compound libraries, decomposing molecules into reusable fragments, assembling novel candidates, and evaluating them through configurable screening pipelines. The platform extends from core cheminformatics into protein target management, molecular docking with AutoDock Vina, and a plugin marketplace for third-party tool integration.
Quick Start¶
# Install dependencies
pip install -r requirements.txt
# Run database migrations
alembic upgrade head
# Seed the fragment library
python scripts/seed_fragments.py
# Start the development server
uvicorn chemlib.main:app --reload
Platform Modules¶
-
Chemical Library
Fragment-based compound management with BRICS decomposition, molecular assembly, and drug-likeness scoring.
-
Protein Targets
Protein target library with structure management, binding site detection, and sequence alignment tools.
-
Structural Biology
Sequence and structural alignment tools for comparing protein targets and identifying conserved binding regions.
-
Virtual Screening
Configurable DAG-based screening pipeline for filtering and ranking compound libraries against targets.
-
Molecular Docking
AutoDock Vina integration with automated pocket detection (Fpocket) and interaction analysis (PLIP).
-
Plugin Marketplace
Extensible architecture for third-party tools, custom scoring functions, and community-contributed plugins.
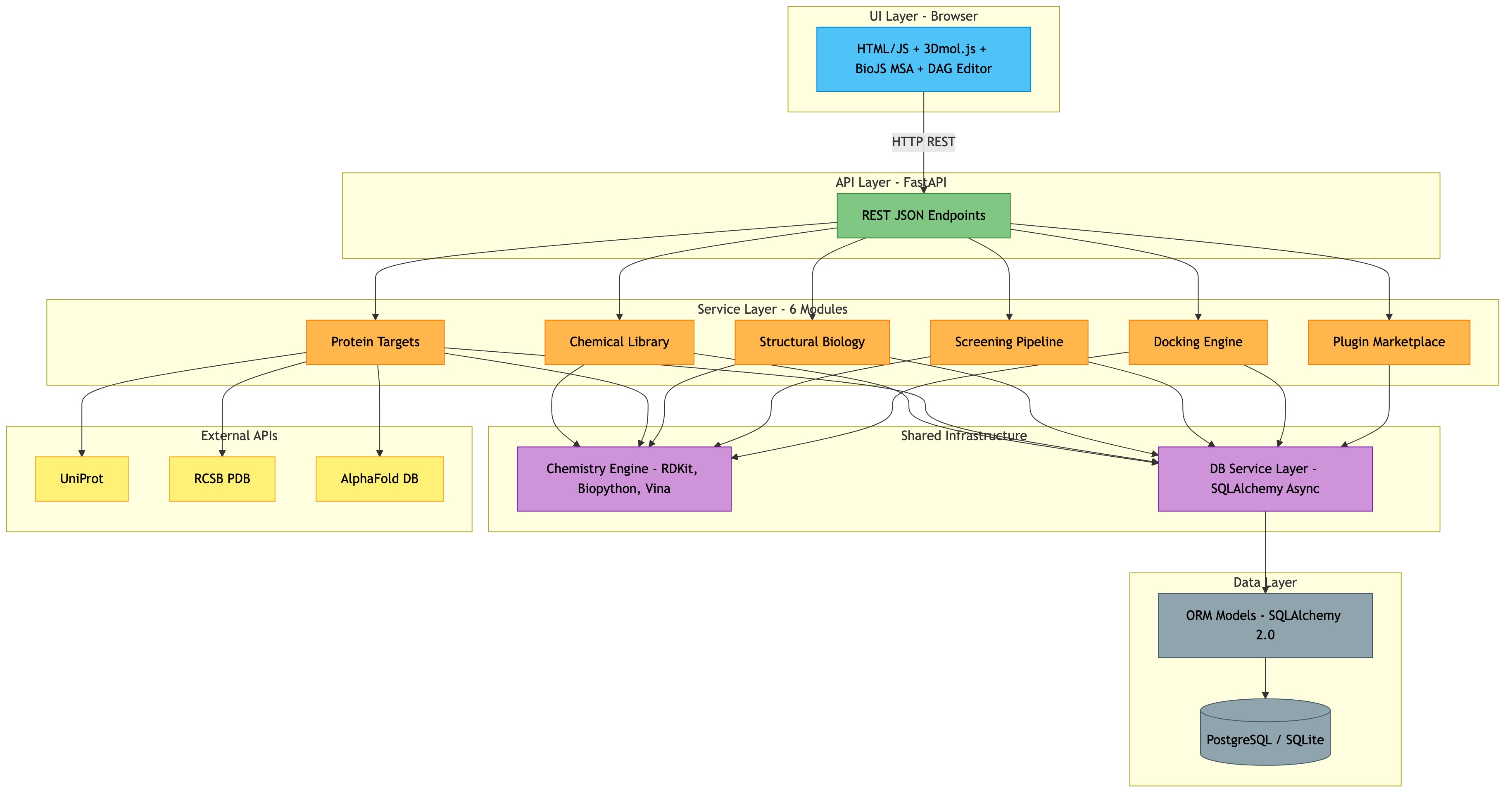
Architecture¶
The platform follows a strict layered architecture: UI, API, Services, Database Service, ORM Models, and Database. All layers communicate through well-defined interfaces, ensuring clean separation of concerns.