Overview
Click diagram to zoom and pan:
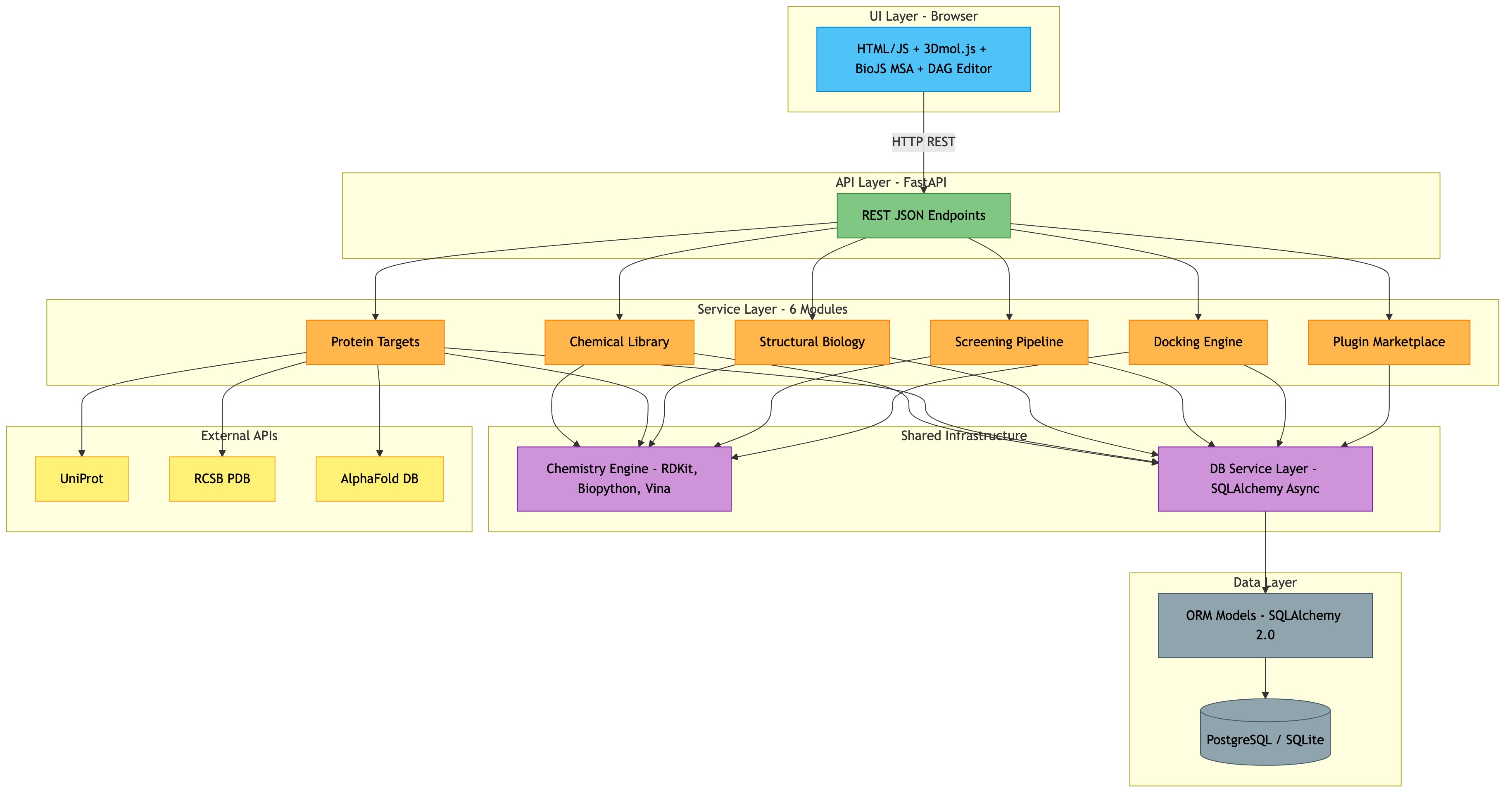
This document defines the phased implementation plan for extending ChemLib from a chemical library tool into a full Computer-Aided Drug Discovery (CADD) platform. The existing system (Phases 1-6) provides compound management, fragment decomposition, BRICS assembly, drug-likeness scoring, and 3D visualization. Phases 7-12 add protein target management, structural biology tools, molecular docking, screening pipelines, and a plugin marketplace.
Prerequisites
Before starting Phase 7, the following must be complete and working:
- Phase 1: Database + ORM models + Alembic migrations
- Phase 2: Chemistry engine (RDKit utilities)
- Phase 3: API layer (FastAPI routes, Pydantic schemas)
- Phase 4: Assembly engine (BRICS fragment joining)
- Phase 5: Scoring and filtering (Lipinski, PAINS, QED, SA Score)
- Phase 6: 3D visualization (3Dmol.js, conformer generation)
All existing tests pass. The application runs with uvicorn chemlib.main:app --reload.
Click diagram to zoom and pan:
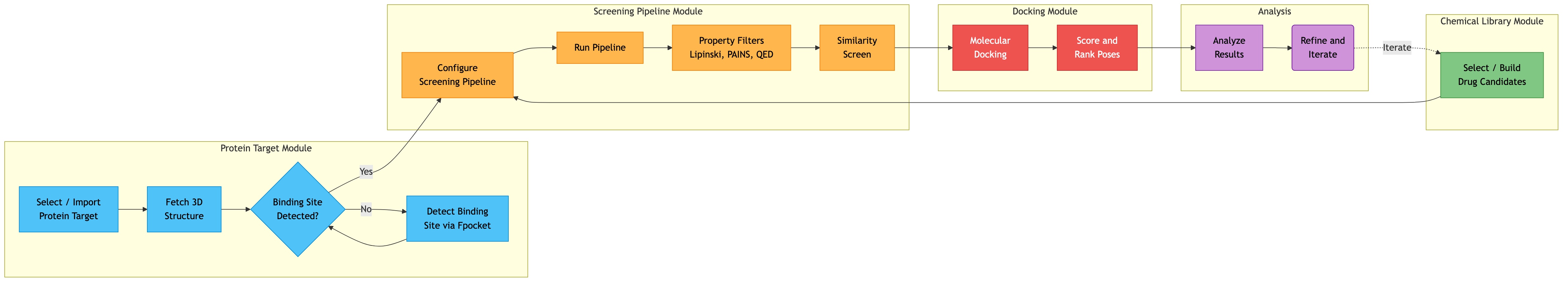
Dependency Graph
Phase 1-6 (existing ChemLib) ─────────────────────────────────────────────┐
│ │
├──────────────┬─────────────────┐ │
▼ ▼ │ │
Phase 7 Phase 8 │ │
Protein Structural │ │
Targets Biology │ │
│ │ │ │
▼ │ │ │
Phase 9 ◀──────────┘ │ │
Binding Site │ │
Detection │ │
│ │ │
▼ │ │
Phase 10 │ │
Docking │ │
Engine │ │
│ │ │
▼ ▼ │
Phase 11 ◀───────────────────────────┘ │
Screening │
Pipeline │
│ │
▼ │
Phase 12 │
Plugin │
Marketplace │
Parallelizable: Phases 7 and 8 can be built simultaneously.
Phase 7: Protein Target Library
Goal: Enable users to import, browse, and manage protein targets and their 3D structures from UniProt, RCSB PDB, and AlphaFold DB.
Design Document: docs/PROTEIN_TARGET_LIBRARY.md
New Dependencies
biopython>=1.83
tmtools>=0.1.0
httpx (already present)
Tasks
7.1 Database Models
| File |
Contents |
chemlib/models/protein.py |
ProteinTarget, ProteinStructure, BindingSite ORM models |
chemlib/models/__init__.py |
Register new models with Base |
- Create all three models with full column definitions per PROTEIN_TARGET_LIBRARY.md
- Add relationships: ProteinTarget → ProteinStructure (1:N), ProteinStructure → BindingSite (1:N)
- Alembic migration:
alembic revision --autogenerate -m "add protein target tables"
- Test: model creation, relationships, cascade delete
7.2 External API Clients
| File |
Contents |
chemlib/bioinformatics/__init__.py |
Package init |
chemlib/bioinformatics/external_apis.py |
UniProtClient, RCSBClient, AlphaFoldClient |
- Implement HTTP clients using
httpx.AsyncClient
- UniProt: fetch entry, search, parse entry
- RCSB: fetch entry info, download PDB, download mmCIF, get UniProt mapping
- AlphaFold: fetch prediction, download PDB
- Test with mocked responses (record real responses as fixtures)
7.3 PDB Parser Utilities
| File |
Contents |
chemlib/bioinformatics/pdb_parser.py |
PDB/mmCIF parsing functions |
parse_pdb_string(), parse_mmcif_string() using Biopythonextract_chains(), extract_ligands(), extract_sequence_from_chain()get_resolution(), get_method()- Test with sample PDB files
7.4 Pydantic Schemas
| File |
Contents |
chemlib/schemas/protein.py |
All Pydantic models for protein targets, structures, binding sites |
ProteinTargetCreate, ProteinTargetResponse, ProteinTargetFilter, ProteinTargetListResponseProteinStructureCreate, ProteinStructureResponse, ProteinStructureDetailResponseBindingSiteCreate, BindingSiteFromLigand, BindingSiteResponse
7.5 DB Service Layer
| File |
Contents |
chemlib/db/service.py |
Add ProteinTargetDBService, ProteinStructureDBService, BindingSiteDBService |
- CRUD operations for all three models
- Specialized queries: list structures for target, list binding sites for structure
7.6 Business Services
| File |
Contents |
chemlib/services/protein_target_service.py |
ProteinTargetService |
chemlib/services/protein_structure_service.py |
ProteinStructureService |
import_from_uniprot(accession): call UniProtClient, parse, storeimport_from_pdb(pdb_id): call RCSBClient, resolve UniProt, storesearch_uniprot(query): proxy search to UniProt APIfetch_from_rcsb(pdb_id, target_id): download PDB, parse metadata, storefetch_from_alphafold(uniprot_id, target_id): download AlphaFold PDB, storeupload_structure(target_id, file_data): validate, parse, store
7.7 API Routes
| File |
Contents |
chemlib/api/targets.py |
/api/targets/ CRUD + fetch endpoints |
chemlib/api/structures.py |
/api/structures/ CRUD + chain/sequence endpoints |
- Register routes in
chemlib/main.py
- All endpoints per PROTEIN_TARGET_LIBRARY.md API section
7.8 UI
| File |
Contents |
chemlib/templates/protein_browser.html |
Protein target list page |
chemlib/templates/protein_detail.html |
Protein detail with 3D viewer |
chemlib/static/js/protein_viewer.js |
3Dmol.js protein viewer component |
- Protein browser: table with search, filter, pagination
- Protein detail: metadata display, structure list, 3D viewer (cartoon mode)
- Viewer: load PDB data, cartoon/surface/stick styles, chain selection
7.9 Tests
| Directory |
Contents |
tests/test_protein/test_models.py |
ORM model tests |
tests/test_protein/test_services.py |
Service tests with mocked APIs |
tests/test_protein/test_api.py |
API endpoint tests |
tests/test_bioinformatics/test_pdb_parser.py |
PDB parsing tests |
tests/test_bioinformatics/test_external_apis.py |
API client tests (mocked) |
Deliverable
- Protein target browser with import from UniProt
- 3D structure viewer (cartoon mode) for PDB structures fetched from RCSB/AlphaFold
- Full CRUD API for targets and structures
Goal: Add sequence alignment (pairwise + MSA), structural alignment (TM-align), and visualization of alignment results.
Design Document: docs/PROTEIN_TARGET_LIBRARY.md (alignment sections)
Can be built in parallel with Phase 7.
New Dependencies
# Python packages
tmtools>=0.1.0 (may already be added in Phase 7)
pymsaviz>=0.4.0
# System binaries (must be on PATH)
mafft # brew install mafft / apt install mafft
Tasks
8.1 Database Models
| File |
Contents |
chemlib/models/alignment.py |
SequenceAlignment, StructuralAlignment ORM models |
- Alembic migration: add alignment tables
- Note:
StructuralAlignment has FK to ProteinStructure — requires Phase 7 tables to exist
8.2 Sequence Alignment Utilities
| File |
Contents |
chemlib/bioinformatics/sequence_tools.py |
pairwise_align_biopython(), multiple_align_mafft(), multiple_align_clustalo() |
- Biopython PairwiseAligner with BLOSUM62
- MAFFT subprocess wrapper (write FASTA temp file, parse output)
- Identity percentage computation
- Test with known sequence pairs
8.3 Structural Alignment Utilities
| File |
Contents |
chemlib/bioinformatics/structural_tools.py |
tm_align(), superimpose_biopython(), apply_transformation() |
- tmtools for TM-align: extract CA coords, run alignment, get rotation/translation
- Biopython Superimposer as fallback
- Apply transformation to generate superposed PDB
- Test with known structure pairs
8.4 Pydantic Schemas
| File |
Contents |
chemlib/schemas/alignment.py |
SequenceAlignmentRequest, SequenceInput, SequenceAlignmentResponse, StructuralAlignmentRequest, StructuralAlignmentResponse |
8.5 Business Services
| File |
Contents |
chemlib/services/alignment_service.py |
AlignmentService |
pairwise_sequence_align(): validate inputs, run alignment, store resultmultiple_sequence_align(): validate inputs, run MAFFT, store resultstructural_align(): load structures, run tmtools/Superimposer, store resultgenerate_alignment_image(): use pyMSAviz to render PNG
8.6 API Routes
| File |
Contents |
chemlib/api/alignments.py |
/api/alignments/ endpoints |
- POST
/api/alignments/sequence — run sequence alignment
- GET
/api/alignments/sequence/{id} — get result
- GET
/api/alignments/sequence/{id}/image — get PNG image
- POST
/api/alignments/structure — run structural alignment
- GET
/api/alignments/structure/{id} — get result
- Register in
chemlib/main.py
8.7 UI
| File |
Contents |
chemlib/templates/alignment_viewer.html |
Alignment results page |
chemlib/static/js/msa_viewer.js |
BioJS MSA Viewer integration |
- Sequence alignment viewer: BioJS MSA Viewer with color schemes (Clustal, Zappo)
- Structural alignment viewer: 3Dmol.js overlay of two structures with different colors
- Display metrics: identity %, score, TM-score, RMSD
- Download buttons: FASTA, PNG image
8.8 Tests
| Directory |
Contents |
tests/test_alignment/test_sequence.py |
Sequence alignment tests |
tests/test_alignment/test_structural.py |
Structural alignment tests (requires tmtools) |
tests/test_alignment/test_api.py |
API endpoint tests |
Deliverable
- Pairwise sequence alignment with Biopython
- Multiple sequence alignment with MAFFT
- Structural alignment with TM-align (tmtools)
- Interactive alignment viewers in the browser
Phase 9: Binding Site Detection & Protein Preparation
Goal: Detect druggable binding pockets using Fpocket, define binding sites from co-crystallized ligands or manually, and prepare proteins for docking.
Depends on: Phase 7 (ProteinStructure, BindingSite models must exist)
New Dependencies
# Python packages
pdbfixer>=1.9
# System binaries
fpocket # brew install fpocket / compile from source
Tasks
9.1 Fpocket Integration
| File |
Contents |
chemlib/bioinformatics/pocket_detection.py |
run_fpocket(), parse_fpocket_info(), parse_pocket_pdb() |
- Write PDB to temp file, run
fpocket -f file.pdb as subprocess
- Parse output: info.txt (scores), pocket PDB files (coordinates)
- Extract center, box size, residues, druggability score, volume per pocket
- Clean up temp files
- Test with a small known protein (mark as integration test, skip if fpocket not installed)
9.2 Protein Preparation
| File |
Contents |
chemlib/bioinformatics/protein_prep.py |
fix_protein(), remove_water(), add_hydrogens() |
- PDBFixer integration for fixing missing atoms/residues
- Remove heterogens and water
- Add hydrogens at specified pH
- Test with broken PDB files
9.3 Binding Site Service
| File |
Contents |
chemlib/services/binding_site_service.py |
BindingSiteService |
detect_pockets(structure_id): run Fpocket, parse, store BindingSite recordsdefine_from_ligand(structure_id, ligand_id, padding): find ligand atoms, compute center/box, find nearby residuesdefine_manual(structure_id, data): store user-defined binding site- CRUD operations for binding sites
9.4 Protein Preparation Service
| File |
Contents |
chemlib/services/protein_prep_service.py |
ProteinPrepService |
prepare_for_docking(structure_id): fix protein, add hydrogens, return fixed PDB
9.5 API Extensions
Add to existing chemlib/api/structures.py:
- POST /api/structures/{id}/detect-pockets — run Fpocket
- POST /api/structures/{id}/binding-sites — define manual
- POST /api/structures/{id}/binding-sites/from-ligand — define from ligand
- GET /api/structures/{id}/binding-sites — list binding sites
- POST /api/structures/{id}/prepare — prepare for docking
9.6 UI Extensions
Update chemlib/templates/protein_detail.html:
- Add binding site list section
- Add "Detect Pockets" button
- 3Dmol.js: show binding site box overlay, residue highlighting
- Binding site detail modal with center/box/residues
9.7 Tests
| Directory |
Contents |
tests/test_protein/test_binding_site.py |
Binding site service tests |
tests/test_bioinformatics/test_pocket_detection.py |
Fpocket tests (integration, skip if binary missing) |
tests/test_bioinformatics/test_protein_prep.py |
PDBFixer tests |
Deliverable
- Fpocket pocket detection with druggability scores
- Binding site definition from ligand or manual
- Protein preparation (fix, add H) for docking
- Binding site visualization in 3D viewer
Phase 10: Docking Engine
Goal: Integrate AutoDock Vina for molecular docking, meeko for ligand preparation, PLIP for interaction analysis, and visualization of docked poses.
Design Document: docs/DOCKING_INTEGRATION.md
Depends on: Phase 9 (binding sites, protein preparation)
New Dependencies
# Python packages
vina>=1.2.5
meeko>=0.5.0
openbabel-wheel>=3.1.0
plip>=2.3.0
Tasks
10.1 Database Models
| File |
Contents |
chemlib/models/docking.py |
DockingRun, DockingResult ORM models |
- Alembic migration: add docking tables
- FKs to: ProteinStructure, BindingSite, Compound, AssembledMolecule
10.2 Docking Utilities
| File |
Contents |
chemlib/docking/__init__.py |
Package init |
chemlib/docking/ligand_prep.py |
smiles_to_pdbqt(), mol_to_pdbqt(), pdbqt_to_pdb(), split_pdbqt_poses() |
chemlib/docking/receptor_prep.py |
pdb_to_pdbqt(), prepare_receptor_full() |
chemlib/docking/vina_runner.py |
dock_ligand(), score_ligand() |
chemlib/docking/interaction_analysis.py |
analyze_complex() |
- meeko for ligand PDBQT conversion
- Open Babel subprocess for receptor PDBQT conversion
- Vina Python API for docking
- PLIP for interaction profiling
- Test each utility independently
10.3 Pydantic Schemas
| File |
Contents |
chemlib/schemas/docking.py |
DockingRunCreate, DockingRunResponse, DockingResultResponse, DockingResultDetailResponse, InteractionAnalysisResponse |
10.4 Business Services
| File |
Contents |
chemlib/services/docking_service.py |
DockingService |
chemlib/services/interaction_service.py |
InteractionService |
prepare_receptor(): PDBFixer + PDBQT conversion, cacheprepare_ligand(): SMILES → 3D → meeko → PDBQTdock(): batch docking — prepare receptor once, dock each liganddock_single(): low-level single ligand dockinganalyze_interactions(): PLIP on protein-ligand complex- Background execution for batch docking
10.5 API Routes
| File |
Contents |
chemlib/api/docking.py |
/api/docking/ endpoints |
- Full endpoint set per DOCKING_INTEGRATION.md
- Background task for batch docking runs
- Register in
chemlib/main.py
10.6 UI
| File |
Contents |
chemlib/templates/docking_viewer.html |
Docking results page |
chemlib/static/js/protein_viewer.js |
Extend with docking pose display |
- 3D viewer: protein (cartoon, gray) + ligand (sticks, green) + binding site (surface)
- Interaction display: H-bond dashes, contact labels
- Results table: ranked by score, interaction counts
- Pose selector dropdown (switch between top N poses)
- Interaction diagram (2D summary)
10.7 Tests
| Directory |
Contents |
tests/test_docking/test_ligand_prep.py |
meeko conversion tests |
tests/test_docking/test_receptor_prep.py |
PDBQT conversion tests |
tests/test_docking/test_vina.py |
Docking tests (integration, requires vina) |
tests/test_docking/test_plip.py |
Interaction analysis tests |
tests/test_docking/test_service.py |
DockingService tests |
tests/test_docking/test_api.py |
API endpoint tests |
Deliverable
- AutoDock Vina docking via API
- Ligand + receptor preparation pipeline
- PLIP interaction analysis
- 3D docking pose viewer with interaction overlay
- Batch docking with progress tracking
Phase 11: Screening Pipeline Engine
Goal: Build the configurable, DAG-based screening pipeline engine with a visual editor, background execution, and results visualization.
Design Document: docs/SCREENING_PIPELINE.md
Depends on: Phase 10 (docking as a filter node) + existing ChemLib (compounds, scoring)
New Dependencies
None — uses existing tools.
Tasks
11.1 Plugin Protocol and Built-in Filters
| File |
Contents |
chemlib/plugins/__init__.py |
Package init |
chemlib/plugins/protocols.py |
FilterPlugin, FilterResult, PipelineContext protocols/dataclasses |
chemlib/plugins/builtin/__init__.py |
Package init |
chemlib/plugins/builtin/property_filters.py |
LipinskiFilter, VeberFilter, GhoseFilter, EganFilter, MueggeFilter, PAINSFilter, BrenkFilter, QEDThresholdFilter, SAScoreFilter, MWRangeFilter, LogPRangeFilter, TPSARangeFilter, HBDMaxFilter, HBAMaxFilter, RotBondsMaxFilter |
chemlib/plugins/builtin/similarity_filters.py |
TanimotoSimilarityFilter, SubstructureMatchFilter, MACCSSimilarityFilter |
chemlib/plugins/builtin/adme_filters.py |
ESOLSolubilityFilter, BBBRuleFilter, hERGRuleFilter, RuleOfThreeFilter |
chemlib/plugins/builtin/docking_filter.py |
VinaDockingFilter |
chemlib/plugins/builtin/external_filters.py |
PLIPInteractionFilter, ADMETlabFilter (stubbed) |
- Each filter implements the full
FilterPlugin protocol
- Each has proper
config_schema (JSON Schema)
- Test each filter independently with known molecules
11.2 Database Models
| File |
Contents |
chemlib/models/pipeline.py |
Pipeline, PipelineRun, PipelineRunResult, FilterPluginRegistry ORM models |
- Alembic migration: add pipeline and plugin tables
- FKs to: ProteinTarget, Compound, AssembledMolecule
11.3 Pydantic Schemas
| File |
Contents |
chemlib/schemas/pipeline.py |
PipelineDefinition, PipelineNode, PipelineEdge, PipelineCreate, PipelineResponse, PipelineRunCreate, PipelineRunResponse, PipelineRunResultResponse, PipelineRunResultFilter, PluginRegistryResponse |
11.4 Pipeline Executor
| File |
Contents |
chemlib/services/pipeline_executor.py |
PipelineExecutor class |
- DAG validation (no cycles)
- Topological sort (Kahn's algorithm)
- Plugin instantiation from registry
- Batch processing with configurable batch size
- Early termination (skip downstream for failed compounds)
- Progress tracking (update PipelineRun status)
- Results storage (PipelineRunResult per compound per node)
- Error handling (catch plugin errors, mark compound as failed, continue)
- Test with a simple 3-node pipeline and mock filters
11.5 Business Services
| File |
Contents |
chemlib/services/pipeline_service.py |
PipelineService |
- CRUD for pipeline definitions
- DAG validation on create/update
- Start run: resolve compound list, create PipelineRun, spawn background executor
- Get run status, results, funnel summary
- Cancel run
11.6 Plugin Seeding Script
| File |
Contents |
scripts/seed_plugins.py |
Register all built-in plugins in the database |
- Called once during setup or on app startup
- Upserts to avoid duplicates
11.7 API Routes
| File |
Contents |
chemlib/api/pipelines.py |
/api/pipelines/ endpoints |
chemlib/api/plugins.py |
/api/plugins/ endpoints |
- Full endpoint set per SCREENING_PIPELINE.md
- Background task for pipeline execution
- Register in
chemlib/main.py
11.8 UI
| File |
Contents |
chemlib/templates/pipeline_builder.html |
Visual DAG editor page |
chemlib/templates/pipeline_results.html |
Run results and funnel visualization |
chemlib/static/js/pipeline_editor.js |
DAG editor (drag-and-drop nodes, connect edges) |
chemlib/static/js/plugin_config_form.js |
JSON Schema → HTML form renderer |
- Pipeline builder: sidebar with filter nodes by category, canvas for DAG, config panel
- Node drag-and-drop, edge drawing, node configuration
- Serialize/deserialize to PipelineDefinition JSON
- Results page: funnel bar chart, filterable results table
- Progress polling during pipeline execution
11.9 Tests
| Directory |
Contents |
tests/test_plugins/test_property_filters.py |
All property filter tests |
tests/test_plugins/test_similarity_filters.py |
Similarity filter tests |
tests/test_plugins/test_adme_filters.py |
ADME filter tests |
tests/test_pipeline/test_executor.py |
Pipeline executor tests |
tests/test_pipeline/test_service.py |
Pipeline service tests |
tests/test_pipeline/test_api.py |
API endpoint tests |
Deliverable
- 25+ built-in filter plugins
- Visual pipeline builder (DAG editor)
- Background pipeline execution with progress tracking
- Funnel visualization of results
Phase 12: Plugin Marketplace
Goal: Formalize the plugin architecture, add entry point discovery, build the marketplace UI, and auto-generate config forms.
Design Document: docs/PLUGIN_MARKETPLACE.md
Depends on: Phase 11 (plugin protocols, registry, pipeline integration)
New Dependencies
None — uses existing infrastructure.
Tasks
12.1 Plugin Registry Service
| File |
Contents |
chemlib/plugins/registry.py |
PluginRegistryService |
discover_and_register_all(): scan built-in + entry points_register_builtin_plugins(): import all classes from chemlib.plugins.builtin.*_register_entrypoint_plugins(): scan chemlib.plugins.filter, chemlib.plugins.docking, chemlib.plugins.adme entry point groupsget_plugin_instance(): instantiate and cache pluginslist_plugins(), get_plugin_info(), set_active()- Integrate with app startup in
chemlib/main.py (call discover_and_register_all on startup)
12.2 Additional Protocol Classes
| File |
Contents |
chemlib/plugins/protocols.py |
Add DockingPlugin, ADMEPlugin, VisualizationPlugin, ExternalServicePlugin |
- Full Protocol definitions per PLUGIN_MARKETPLACE.md
DockingPluginResult, ADMEPrediction dataclasses
12.3 API Extensions
Extend chemlib/api/plugins.py:
- PUT /api/plugins/{name}/active — enable/disable
- POST /api/plugins/refresh — re-scan and register
12.4 Marketplace UI
| File |
Contents |
chemlib/templates/plugin_marketplace.html |
Plugin browser page |
chemlib/static/js/plugin_config_form.js |
Enhance JSON Schema form renderer |
- Browse plugins by category (card layout)
- Each card: name, description, category, estimated time, active status
- "Configure" button: opens modal with auto-generated form
- "Use in Pipeline" button: redirects to pipeline builder with plugin pre-selected
- Category filter, search bar
- Admin: enable/disable toggle
12.5 App Startup Integration
Update chemlib/main.py:
@app.on_event("startup")
async def register_plugins():
async with get_db_session() as db:
registry = PluginRegistryService()
await registry.discover_and_register_all(db)
12.6 Entry Point Documentation
Create example for third-party plugin developers:
- How to create a plugin package
- How to implement FilterPlugin protocol
- How to define entry points in pyproject.toml
- How to test plugin compliance
12.7 Tests
| Directory |
Contents |
tests/test_plugins/test_registry.py |
Plugin discovery and registration tests |
tests/test_plugins/test_protocol_compliance.py |
Verify all built-in plugins satisfy Protocol |
tests/test_plugins/test_api.py |
Plugin API endpoint tests |
Deliverable
- Plugin registry with automatic discovery
- Marketplace UI for browsing and configuring plugins
- Entry point-based plugin installation support
- All built-in plugins registered and available
Phase Summary Table
| Phase |
Name |
New Python Deps |
System Binaries |
Key Deliverable |
Est. Effort |
| 7 |
Protein Target Library |
biopython, tmtools |
— |
Protein browser + 3D viewer |
Medium |
| 8 |
Structural Biology |
pymsaviz |
mafft |
Alignment tools + visualization |
Medium |
| 9 |
Binding Site Detection |
pdbfixer |
fpocket |
Pocket detection + protein prep |
Small |
| 10 |
Docking Engine |
vina, meeko, openbabel-wheel, plip |
— |
Molecular docking pipeline |
Large |
| 11 |
Screening Pipeline |
— |
— |
Visual pipeline builder + executor |
Large |
| 12 |
Plugin Marketplace |
— |
— |
Extensible plugin architecture |
Medium |
Configuration Updates
chemlib/config.py Additions
class Settings(BaseSettings):
# ... existing settings ...
# Phase 7: External APIs
UNIPROT_API_BASE: str = "https://rest.uniprot.org"
RCSB_API_BASE: str = "https://data.rcsb.org"
RCSB_FILES_BASE: str = "https://files.rcsb.org"
ALPHAFOLD_API_BASE: str = "https://alphafold.ebi.ac.uk/api"
EXTERNAL_API_TIMEOUT: int = 30 # seconds
# Phase 8: Alignment tools
MAFFT_BINARY: str = "mafft"
CLUSTALO_BINARY: str = "clustalo"
FOLDSEEK_BINARY: str = "foldseek"
# Phase 9: Pocket detection
FPOCKET_BINARY: str = "fpocket"
DEFAULT_POCKET_MIN_DRUGGABILITY: float = 0.2
DEFAULT_BINDING_SITE_PADDING: float = 5.0 # Angstroms
# Phase 10: Docking
VINA_EXHAUSTIVENESS: int = 32
VINA_NUM_POSES: int = 10
VINA_ENERGY_RANGE: float = 3.0
DOCKING_DEFAULT_PH: float = 7.0
# Phase 11: Pipeline
PIPELINE_BATCH_SIZE: int = 100
PIPELINE_MAX_COMPOUNDS: int = 100_000
PIPELINE_POLL_INTERVAL: int = 3 # seconds (for UI polling)
requirements.txt Additions
# Phase 7
biopython>=1.83
tmtools>=0.1.0
# Phase 8
pymsaviz>=0.4.0
# Phase 9
pdbfixer>=1.9
# Phase 10
vina>=1.2.5
meeko>=0.5.0
openbabel-wheel>=3.1.0
plip>=2.3.0
Migration Strategy
Each phase creates its own Alembic migration. Migrations are ordered and can be applied incrementally:
# Phase 7
alembic revision --autogenerate -m "add protein target, structure, binding site tables"
alembic upgrade head
# Phase 8
alembic revision --autogenerate -m "add sequence and structural alignment tables"
alembic upgrade head
# Phase 10
alembic revision --autogenerate -m "add docking run and docking result tables"
alembic upgrade head
# Phase 11
alembic revision --autogenerate -m "add pipeline, pipeline run, pipeline result, filter plugin registry tables"
alembic upgrade head
Important: Phase 9 does not need its own migration because BindingSite is created in Phase 7's migration (it belongs to the protein module). Phase 12 does not need new tables — it uses the FilterPluginRegistry from Phase 11.
Router Registration Order
Update chemlib/main.py as phases are completed:
# chemlib/main.py
from fastapi import FastAPI
from chemlib.api import compounds, fragments, assembly, visualization, scoring # Existing
from chemlib.api import targets, structures # Phase 7
from chemlib.api import alignments # Phase 8
# Phase 9 endpoints are in structures.py (already registered)
from chemlib.api import docking # Phase 10
from chemlib.api import pipelines, plugins # Phase 11-12
app = FastAPI(title="ChemLib Drug Discovery Platform")
# Existing
app.include_router(compounds.router)
app.include_router(fragments.router)
app.include_router(assembly.router)
app.include_router(visualization.router)
app.include_router(scoring.router)
# Phase 7
app.include_router(targets.router)
app.include_router(structures.router)
# Phase 8
app.include_router(alignments.router)
# Phase 10
app.include_router(docking.router)
# Phase 11-12
app.include_router(pipelines.router)
app.include_router(plugins.router)
Quality Checklist (Per Phase)
Before marking a phase as complete, verify:
- [ ] All ORM models match the design document schemas exactly
- [ ] Alembic migration applies cleanly on fresh DB and existing DB
- [ ] All service methods have proper error handling (custom exceptions)
- [ ] All API endpoints have Pydantic request/response validation
- [ ] All endpoints are documented in OpenAPI (auto-generated from FastAPI)
- [ ] External API calls handle timeouts, rate limits, and error responses
- [ ] Subprocess calls (Fpocket, MAFFT, Open Babel) handle missing binaries gracefully
- [ ] Background tasks properly update status on success and failure
- [ ] UI pages are functional (load data, display results, handle errors)
- [ ] Unit tests pass for all new utility functions
- [ ] Integration tests pass for services
- [ ] E2E tests pass for API endpoints
- [ ] No regressions in existing tests
- [ ]
CLAUDE.md is updated with new modules and commands

