Database Design — ChemLib¶
Overview¶
The database stores chemical compounds, fragments, assembled molecules, and all associated structural representations (1D, 2D, 3D). SQLAlchemy 2.0 with async support serves as the ORM. Alembic manages all schema migrations.
Database Choice¶
- Development: SQLite via
aiosqlite(zero setup, file-based) - Production: PostgreSQL via
asyncpg(with optional RDKit cartridge for server-side chemical queries) - The application must work with both backends. Avoid PostgreSQL-only features in the ORM; use them only in optional optimized query paths.
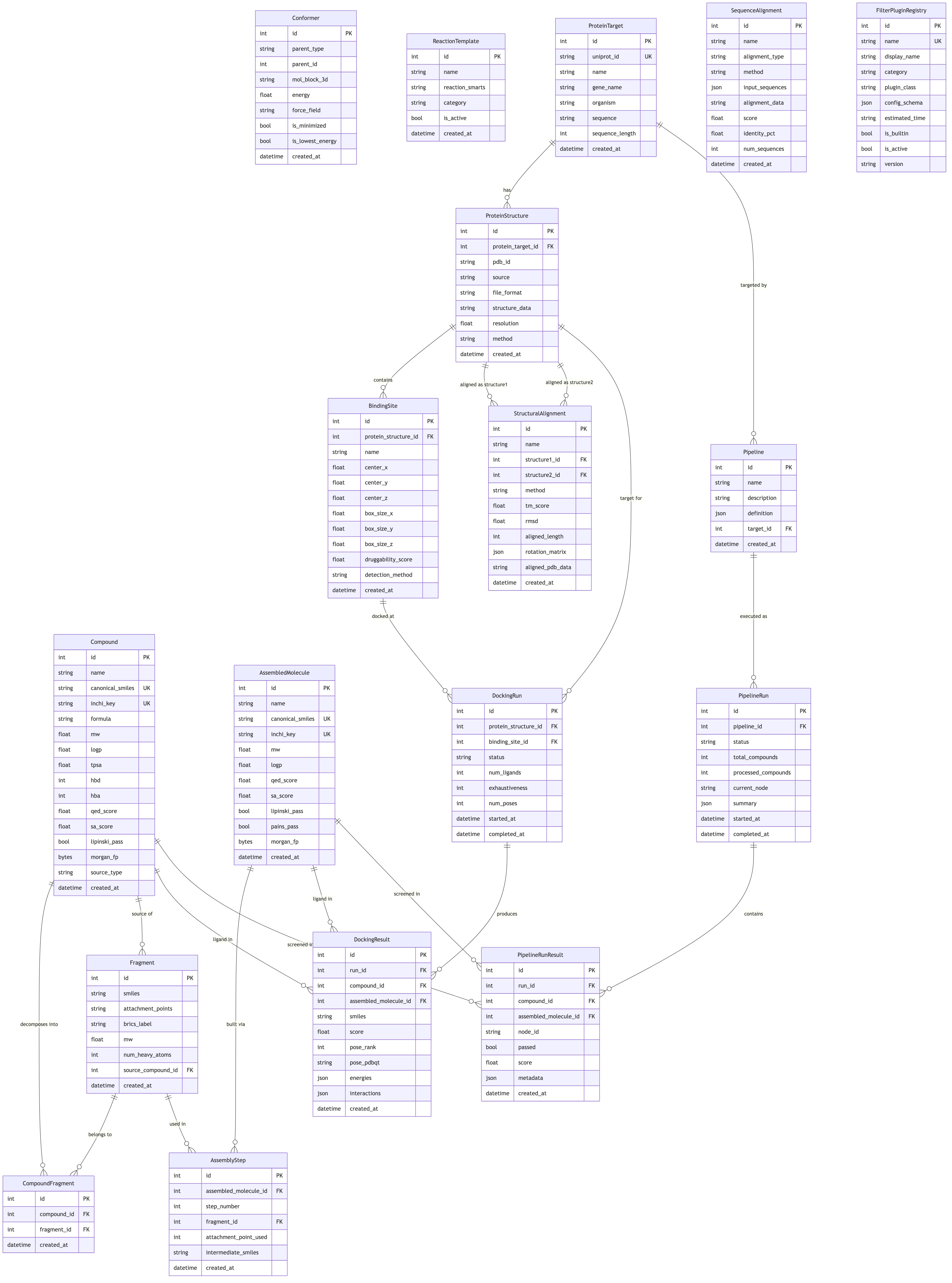
Entity-Relationship Diagram¶
Click diagram to zoom and pan:
┌──────────────────┐ ┌──────────────────────┐
│ Compound │ │ Fragment │
├──────────────────┤ ├──────────────────────┤
│ id (PK) │ │ id (PK) │
│ name │ │ smiles (canonical) │
│ canonical_smiles │ │ attachment_points │ (JSON: list of dummy atom labels)
│ inchi │ │ brics_label │
│ inchi_key (UQ) │ │ mol_block_2d │
│ mol_block_2d │ │ mw │
│ mol_block_3d │ │ num_heavy_atoms │
│ mw │ │ source_compound_id │──FK──▶ Compound.id
│ logp │ │ created_at │
│ tpsa │ │ updated_at │
│ hbd │ └──────────────────────┘
│ hba │
│ num_rotatable │ ┌──────────────────────┐
│ num_rings │ │ Conformer │
│ qed_score │ ├──────────────────────┤
│ sa_score │ │ id (PK) │
│ lipinski_pass │ │ parent_type │ (compound | assembled)
│ morgan_fp │ │ parent_id │
│ source_type │ │ mol_block_3d │ (MOL block with 3D coords)
│ created_at │ │ energy │ (kcal/mol after minimization)
│ updated_at │ │ force_field │ (MMFF94 | UFF)
│ is_assembled │ │ is_minimized │
│ formula │ │ is_lowest_energy │
│ smiles_depiction │ │ rmsd │
└──────────────────┘ │ created_at │
│ └──────────────────────┘
│
│ (decomposition)
▼
┌──────────────────────────────────┐
│ CompoundFragment (junction) │
├──────────────────────────────────┤
│ id (PK) │
│ compound_id (FK) │──▶ Compound.id
│ fragment_id (FK) │──▶ Fragment.id
│ created_at │
└──────────────────────────────────┘
┌──────────────────────────────┐
│ AssembledMolecule │
├──────────────────────────────┤
│ id (PK) │
│ name │
│ canonical_smiles │
│ inchi │
│ inchi_key (UQ) │
│ mol_block_2d │
│ mw │
│ logp │
│ tpsa │
│ hbd │
│ hba │
│ num_rotatable │
│ qed_score │
│ sa_score │
│ lipinski_pass │
│ pains_pass │
│ morgan_fp │
│ formula │
│ created_at │
│ updated_at │
└──────────────────────────────┘
│
│
▼
┌──────────────────────────────┐
│ AssemblyStep │
├──────────────────────────────┤
│ id (PK) │
│ assembled_molecule_id (FK) │──▶ AssembledMolecule.id
│ step_number │
│ fragment_id (FK) │──▶ Fragment.id
│ attachment_point_used │ (which dummy atom label)
│ target_attachment_point │ (attachment point on growing molecule)
│ intermediate_smiles │ (SMILES after this step)
│ created_at │
└──────────────────────────────┘
┌──────────────────────────────┐
│ ReactionTemplate │
├──────────────────────────────┤
│ id (PK) │
│ name │
│ description │
│ reaction_smarts │ (e.g., '[C:1](=O)O.[N:2]>>[C:1](=O)[N:2]')
│ category │ (amide, ester, C-C, etc.)
│ is_active │
│ created_at │
└──────────────────────────────┘
SQLAlchemy ORM Models¶
Base Model¶
# chemlib/models/base.py
from datetime import datetime
from sqlalchemy import func
from sqlalchemy.orm import DeclarativeBase, Mapped, mapped_column
class Base(DeclarativeBase):
pass
class TimestampMixin:
created_at: Mapped[datetime] = mapped_column(default=func.now())
updated_at: Mapped[datetime] = mapped_column(default=func.now(), onupdate=func.now())
Compound Model¶
# chemlib/models/compound.py
class Compound(Base, TimestampMixin):
__tablename__ = "compounds"
id: Mapped[int] = mapped_column(primary_key=True)
name: Mapped[str | None] = mapped_column(String(255))
canonical_smiles: Mapped[str] = mapped_column(Text, unique=True, index=True)
inchi: Mapped[str | None] = mapped_column(Text)
inchi_key: Mapped[str | None] = mapped_column(String(27), unique=True, index=True)
mol_block_2d: Mapped[str | None] = mapped_column(Text) # MOL block with 2D coords
mw: Mapped[float | None]
logp: Mapped[float | None]
tpsa: Mapped[float | None]
hbd: Mapped[int | None] # H-bond donors
hba: Mapped[int | None] # H-bond acceptors
num_rotatable: Mapped[int | None]
num_rings: Mapped[int | None]
qed_score: Mapped[float | None]
sa_score: Mapped[float | None]
lipinski_pass: Mapped[bool | None]
morgan_fp: Mapped[bytes | None] = mapped_column(LargeBinary) # serialized bit vector
formula: Mapped[str | None] = mapped_column(String(255))
source_type: Mapped[str | None] = mapped_column(String(50)) # 'imported', 'assembled'
# Relationships
fragments: Mapped[list["CompoundFragment"]] = relationship(back_populates="compound")
conformers: Mapped[list["Conformer"]] = relationship(...)
Fragment Model¶
# chemlib/models/compound.py
class Fragment(Base, TimestampMixin):
__tablename__ = "fragments"
id: Mapped[int] = mapped_column(primary_key=True)
smiles: Mapped[str] = mapped_column(Text, index=True) # with dummy atoms [1*], [3*], etc.
attachment_points: Mapped[str] = mapped_column(Text) # JSON: e.g., '[1, 3]' (BRICS labels)
brics_label: Mapped[str | None] = mapped_column(String(50))
mol_block_2d: Mapped[str | None] = mapped_column(Text)
mw: Mapped[float | None]
num_heavy_atoms: Mapped[int | None]
source_compound_id: Mapped[int | None] = mapped_column(ForeignKey("compounds.id"))
# Relationships
source_compound: Mapped["Compound | None"] = relationship()
compound_links: Mapped[list["CompoundFragment"]] = relationship(back_populates="fragment")
Key Design Decisions¶
-
Canonical SMILES as unique key: Every compound gets a canonical SMILES via
Chem.MolToSmiles(mol, canonical=True). This prevents duplicate entries. -
InChIKey for cross-referencing: The 27-character InChIKey allows fast lookup and cross-database matching. Indexed for performance.
-
2D MOL blocks stored as Text: MOL V2000/V3000 blocks are stored as text columns. They contain 2D coordinates for depiction.
-
3D coordinates in Conformer table: 3D structures are stored separately because a molecule can have many conformers. Each conformer has its own MOL block with 3D coordinates, energy value, and minimization status.
-
Morgan fingerprints as binary: Serialized
ExplicitBitVectstored asLargeBinary. For PostgreSQL production, consider the RDKit cartridge's nativebfptype instead. -
Fragment attachment points as JSON: A JSON string like
[1, 3, 16]listing BRICS dummy atom labels. This makes it easy to query for compatible fragments. -
Assembly provenance:
AssemblySteprecords every fragment addition, enabling full replay of how a molecule was built. -
Polymorphic conformers: The
Conformertable usesparent_type+parent_idto reference either aCompoundorAssembledMolecule. This avoids duplicating the conformer table.
Indexes¶
| Table | Column(s) | Type | Purpose |
|---|---|---|---|
| compounds | canonical_smiles | UNIQUE | Exact SMILES lookup, deduplication |
| compounds | inchi_key | UNIQUE | Cross-database matching |
| compounds | mw | BTREE | Range queries (MW filter) |
| compounds | logp | BTREE | Range queries (LogP filter) |
| fragments | smiles | INDEX | Fragment lookup |
| fragments | source_compound_id | INDEX | Find fragments from a compound |
| assembled_molecules | canonical_smiles | UNIQUE | Deduplication |
| assembled_molecules | inchi_key | UNIQUE | Cross-database matching |
| conformers | parent_type, parent_id | INDEX | Find conformers for a molecule |
| conformers | is_lowest_energy | INDEX | Quick access to best conformer |
| assembly_steps | assembled_molecule_id | INDEX | Rebuild assembly history |
Migration Strategy (Alembic)¶
Setup¶
Configuration (alembic/env.py)¶
- Import
Base.metadatafromchemlib.models.base - Configure
target_metadata = Base.metadata - Support both sync (SQLite) and async (PostgreSQL) engines
- Use
render_as_batch=Truefor SQLite ALTER TABLE support
Initial Migration¶
Rules¶
- Every schema change requires a migration — no manual DDL
- Migration messages should be descriptive:
"add_sa_score_to_compounds" - Always review autogenerated migrations before applying
- Downgrade functions must be implemented for reversibility
- Test migrations against both SQLite and PostgreSQL
DB Service Layer¶
The DB service layer (chemlib/db/service.py) provides all CRUD operations. No other layer should construct SQLAlchemy queries.
Generic CRUD Base¶
class CRUDBase(Generic[ModelType]):
def __init__(self, model: type[ModelType]):
self.model = model
async def get(self, db: AsyncSession, id: int) -> ModelType | None
async def get_multi(self, db: AsyncSession, skip: int = 0, limit: int = 100) -> list[ModelType]
async def create(self, db: AsyncSession, obj_in: dict) -> ModelType
async def update(self, db: AsyncSession, db_obj: ModelType, obj_in: dict) -> ModelType
async def delete(self, db: AsyncSession, id: int) -> ModelType | None
Specialized Services¶
class CompoundDBService(CRUDBase[Compound]):
async def get_by_smiles(self, db, smiles: str) -> Compound | None
async def get_by_inchi_key(self, db, inchi_key: str) -> Compound | None
async def filter_by_properties(self, db, mw_range, logp_range, ...) -> list[Compound]
async def search_similar(self, db, fp: bytes, threshold: float) -> list[Compound]
class FragmentDBService(CRUDBase[Fragment]):
async def get_by_attachment_points(self, db, labels: list[int]) -> list[Fragment]
async def get_compatible_fragments(self, db, brics_label: int) -> list[Fragment]
class AssemblyDBService(CRUDBase[AssembledMolecule]):
async def get_with_steps(self, db, id: int) -> AssembledMolecule
async def get_with_conformers(self, db, id: int) -> AssembledMolecule
class ConformerDBService(CRUDBase[Conformer]):
async def get_lowest_energy(self, db, parent_type, parent_id) -> Conformer | None
async def get_all_for_molecule(self, db, parent_type, parent_id) -> list[Conformer]